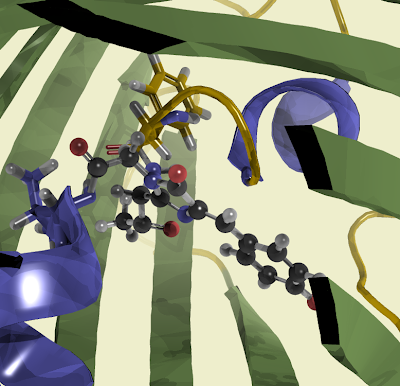
Ribonoclease A is a sturdy little molecule with 124 amino acids. Its four disulfide bridges (yellow in the picture) make it very stable and comparably easy to isolate. Just 124 amino acids also makes it nice for computations. The pictures were made with VMD which makes nice pictures and is also a pretty versatile tool that reads tons of file formats.

The active group of Ribonuclease A comes from two Histidine (black) residues that help hydrolizing the phosphate in the nucleic acid. Lysine (orange) provides the positive charge to interact with negatively charged phosphate groups.
 The first thing you can do with a molecular structure is an energy minimization. These are the Cα traces of the crystal structure (yellow) and the vacuum optimized structure (red). They are in fact quite different even though the red structure is just the next local minimum.
The first thing you can do with a molecular structure is an energy minimization. These are the Cα traces of the crystal structure (yellow) and the vacuum optimized structure (red). They are in fact quite different even though the red structure is just the next local minimum. The major problem with this optimization is that I did not include solvent effects. The electrostatic interaction energy (in the macroscopic limit) is given by
The major problem with this optimization is that I did not include solvent effects. The electrostatic interaction energy (in the macroscopic limit) is given byAnother thing you can do with structures is normal mode analysis. That means computing the eigenvalues/eigenvectors of the mass weighted energy Hessian matrix which correspond to the frequencies/motions of the normal modes. Ribonuclease A is small for a protein but it still has 1856 atoms. So we have 3x1856=5568 degrees of freedom. We have to compute, store and process about 30 million matrix elements. This seems much but these days most regular computers will be able to do this in less than half an hour. 30 million matrix elements corresponds to something like 300 MByte, nothing too crazy. [2]
These are the three lowest frequency modes (aside from translation and rotation) of Ribonuclease A. They are large scale backbone motions. A weak force constant and a high effective mass causes them to have wave numbers below 10/cm.



[1] Not the kind of lab where you need a lab coat.
[2] Actually if you are just interested in a few eigenvalues and -vectors, you can go much higher. For example multi-reference configuration interaction with Columbus (which is a great program by the way....): finding the first 4 states out of a 8569890 x 8569890 matrix was possible within about 17 hours. In this case you don't store trillions (7E13) of matrix elements and do a diagonalization. You use the iterative Davidson procedure which will find an eigenvector close to a starting guess.