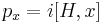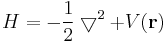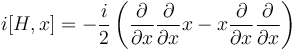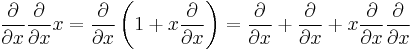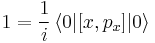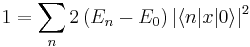W. Zeng, C. Zhong, H. Bronstein, F. Plasser
Understanding and Tuning Singlet-Triplet (S1/T1) Energy Gaps in Planar Organic Chromophores
Angew Chemie Int Ed in print (2025), DOI: 10.1002/anie.202502485
Showing posts with label photochemistry. Show all posts
Showing posts with label photochemistry. Show all posts
Monday, 24 March 2025
Singlet-Triplet Energy Gaps in Planar Organic Chromophores
A new framework for tuning singlet-triplet (S1/T1) gaps based on electrostatic interactions of the transition density is developed. Three widely applicable design rules for maximizing the singlet-triplet gap are derived: minimizing the number of electrons, localizing the excitation, and optimizing through-space interactions.
Saturday, 2 May 2020
De-excitations
If the electronic ground state of a molecule has double-excitation character and the excited state only single-excitation character, then you can view the molecule as being "de-excited." Electrons are taken from a higher lying orbital and moved into a lower lying orbital. A recent paper tries to formalise this idea by computing an expectation value of the particle-hole permutation operator. We take the two-body exciton wavefunction, switch the electron and hole and see how much it resembles the original wavefunction. If the hole resides purely in the occupied orbitals and the electron purely in the virtual orbitals, this has to yield zero. But with a correlated ground state it does not vanish.
The striking thing is that this expectation value of the particle-hole permutation operator seems to agree between TDDFT and wavefunction based methods in, both, magnitude and sign. This means that de-excitations are a "real thing" rather than just an artifact of TDDFT - not physically observable but a well-defined property of the wavefunction. For more on this, see PCCP 2020, 22, 6058.
The striking thing is that this expectation value of the particle-hole permutation operator seems to agree between TDDFT and wavefunction based methods in, both, magnitude and sign. This means that de-excitations are a "real thing" rather than just an artifact of TDDFT - not physically observable but a well-defined property of the wavefunction. For more on this, see PCCP 2020, 22, 6058.
Wednesday, 22 January 2020
Understanding excitation energies beyond the MO picture
Is it possible to get an intuitive understanding of electronic excitation energies that truly goes beyond the MO picture? This is what we are discussing in our newest preprint: Toward an Understanding of Electronic Excitation Energies Beyond the Molecular Orbital Picture. This is basically a sequel to my previous post HOMO-LUMO gaps and excitation energies, which seems to consistently attract visitors to this blog. In fact the paper started out as a short sequel and then it turned into 19 dense pages. Well, I hope it is worth the effort.
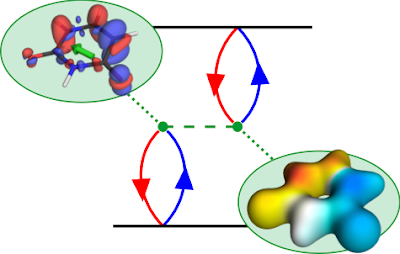
The TOC shows a diagrammatic representation of the exchange repulsion, which is responsible for the difference in energies between singlet and triplet states. The red and blue lines refer to the hole and electron, respectively. The dotted green line is the Coulomb interaction. The diagram is read in the following way: The hole and electron come together on the bra (bottom) and ket (top) side forming the transition density. They interact with each other via the Coulomb interaction. The resulting term can be interpreted as the Coulomb repulsion of the transition density with respect to itself. To represent this, we show the transition density and its electrostatic potential (ESP). The exchange repulsion is now simply an overlap between density and ESP.
The state shown is the first ππ* state of uracil. A closer look at the transition density (upper left) shows the expected π contributions. But why are there also σ contributions? It turns out that the pure ππ* state would have an excessively high exchange repulsion. That is why σ contributions are mixed in to lower the energy. These σ contributions lower the transition moment (shown in green) and, thus, have a direct experimentally observable consequence. They also mean that any description of the state in terms of only n and π orbitals is insufficient, which explains the problems of CASSCF in describing these sorts of states - called ionic states in the valence-bond description.
The TOC shows a diagrammatic representation of the exchange repulsion, which is responsible for the difference in energies between singlet and triplet states. The red and blue lines refer to the hole and electron, respectively. The dotted green line is the Coulomb interaction. The diagram is read in the following way: The hole and electron come together on the bra (bottom) and ket (top) side forming the transition density. They interact with each other via the Coulomb interaction. The resulting term can be interpreted as the Coulomb repulsion of the transition density with respect to itself. To represent this, we show the transition density and its electrostatic potential (ESP). The exchange repulsion is now simply an overlap between density and ESP.
The state shown is the first ππ* state of uracil. A closer look at the transition density (upper left) shows the expected π contributions. But why are there also σ contributions? It turns out that the pure ππ* state would have an excessively high exchange repulsion. That is why σ contributions are mixed in to lower the energy. These σ contributions lower the transition moment (shown in green) and, thus, have a direct experimentally observable consequence. They also mean that any description of the state in terms of only n and π orbitals is insufficient, which explains the problems of CASSCF in describing these sorts of states - called ionic states in the valence-bond description.
Friday, 23 June 2017
Excited State Delocalization in DNA
One persistent topic in photobiology has been the question of how many bases are involved in the absorption of a photon of UV light, or in other words how delocalized the absorbing states are. There have been estimates ranging from completely localized monomeric states to delocalization over the whole helix. The question is challenging to study experimentally because the delocalization of the wavefunction is a quantum phenomenon without a direct experimentally observable counterpart. It is difficult to study from a computational point of view because of the extended system size, environmental interactions, and structural disorder. Computational studies using exciton models could not include structural disorder well. Explicit quantum chemistry studies had troubles because of smaller QM regions.
That is why we thought there was need for another more extended study. We used a QM region of eight nucleobases, which could be treated by TDDFT thanks to the GPU based TeraChem code, we did extended sampling by molecular dynamics using the Amber code (also GPU based), and we connected the two by a QM/MM scheme. In total we computed 6000 excited states. To analyze these systematically, we used the TheoDORE code. The results are shown in a new article "Electronic delocalization, charge transfer and hypochromism in the UV absorption spectrum of polyadenine unravelled by multiscale computations and quantitative wavefunction analysis" that just appeared in Chemical Science.
The main results of the study are:
That is why we thought there was need for another more extended study. We used a QM region of eight nucleobases, which could be treated by TDDFT thanks to the GPU based TeraChem code, we did extended sampling by molecular dynamics using the Amber code (also GPU based), and we connected the two by a QM/MM scheme. In total we computed 6000 excited states. To analyze these systematically, we used the TheoDORE code. The results are shown in a new article "Electronic delocalization, charge transfer and hypochromism in the UV absorption spectrum of polyadenine unravelled by multiscale computations and quantitative wavefunction analysis" that just appeared in Chemical Science.
The main results of the study are:
- Photon absorption occurs predominantly through a collective excitation of two neighboring nucleobases.
- Full charge transfer (CT) states are only present at higher energies but states with non-neglible CT admixture account for about 50% of the spectral intensity.
- The experimentally observed hypochromism occurs through perturbed monomer states rather than excitonic or CT interactions.
Thursday, 5 January 2017
Simulating Light-Induced Processes in DNA
We planned to write a short perspective of our experience of simulating UV excitations in DNA but ended up with a quite comprehensive article: "Challenges in Simulating Light-Induced Processes in DNA", which just appeared in the journal Molecules. The aim of this work was to illustrate the different tasks that are involved in the simulation of DNA and its components: (i) quantum chemistry, (ii) description of the excitation processes, (iii) nonadiabatic dynamics, (iv) comparison to experiment, and (v) analysis of the results. In all these cases significant challenges can occur, and a wide range of methods to tackle these have been developed by numerous researchers. For someone entering the field or even for active researchers, it is sometimes difficult to keep all this in mind. We hope that this new article will be provide a useful summary of the work that has been done.
Friday, 7 March 2014
UV Excited Single- and Double-Stranded DNA
There is a new paper out by us: "Electronic Excitation Processes in Single-Strand and Double-Strand DNA: A Computational Approach" in Topics in Current Chemistry. I made this figure to show all the possible processes happening:
Which ones of these pathways are the important ones is not known yet. The problem is that every research group involved has their own convictions and it is not quite clear who is correct. In this paper we are of course not able to solve the problem, but at least we discuss the different computational methods applied to help make it more clear why the results by different groups are different.
My own contribution to the debate is the hypothesis of an exciplex with strong geometric distortions, small intermolecular separations, and strong orbital interactions that we described in this paper. Some other groups have obtained similar results: In particular Spiridoula Matsika did lots of work (e.g. this paper) where she invokes a bonded exciplex model. And a Chinese group who were the first to obtain these types of results, using semi-empirical calculations.
- Monomer-like decay, which is also observed in the gas phase
- Delocalization and excitation energy transfer (with a focus on electron dynamics)
- Proton transfer between the strands, possibly leading to deactivation
- Electron transfer leading to charge transfer states
- Excimers, which may constitute stable trapping sites
- And finally the thing we are trying to avoid: photoproduct formation, which can lead to damage of the DNA
Which ones of these pathways are the important ones is not known yet. The problem is that every research group involved has their own convictions and it is not quite clear who is correct. In this paper we are of course not able to solve the problem, but at least we discuss the different computational methods applied to help make it more clear why the results by different groups are different.
My own contribution to the debate is the hypothesis of an exciplex with strong geometric distortions, small intermolecular separations, and strong orbital interactions that we described in this paper. Some other groups have obtained similar results: In particular Spiridoula Matsika did lots of work (e.g. this paper) where she invokes a bonded exciplex model. And a Chinese group who were the first to obtain these types of results, using semi-empirical calculations.
Tuesday, 17 December 2013
Oscillator strengths
What is the physical significance of the oscillator strength? Following Werner Kuhn's arguments (e.g. in this paper), it marks the number of electrons oscillating per spatial dimension during an electronic transition. The sum over the oscillator strengths of all the excited states amounts to the number electrons, which is the essence of the Thomas-Reiche-Kuhn sum rule. In other words, the oscillator strength counts how much of the total oscillating potential is used for a specific transition.
This interpretation explains for example the linear relationship of the oscillator strength of the lowest excited state with system size in the case of some conjugated organic polymers (see e.g. this paper): If there are more electrons available to oscillate, then the transition strength increases.
The oscillator strength fij between two non-degenerate states i and j is defined (in atomic units) as two thirds of the squared transition dipole moment multiplied by the energy gap
The Thomas-Reiche-Kuhn sum rule now states that the sum over the oscillator strengths from one state i to all possible other states is equal to the number of electrons in the system, i.e.
In particular, if we consider excitations from the ground state, then all oscillator strengths are positive. Which means that the oscillator strengths can in fact be viewed as a partitioning of the number of electrons.
The derivation of this sum rule starts by realizing that the momentum operator with respect to any spatial coordinate x of any particle (e.g. x=y2) is given as the commutator of the Hamiltonian with this coordinate
This follows whenever H is of the form
where clearly the derivatives with respect to x are the only part, which does not commute with x itself
By applying the product rule twice, the first term of this expression becomes
And in summary
The remaining proof follows what is shown here (sorry that I am switching the notation, but I copy-and-pasted a little bit ...). First one realizes that the commutator of x and px is equal to i
Then one expands the commutators and inserts a resolution of the identity over the eigenstates of the Hamiltonian
Insert the above expression for px
The commutators are evaluated by letting H act either on the bra or the ket, which results in a multiplication with the respective eigenvalue. And after summing together the equivalent terms one obtains
The actual r vector was composed of 3N individual electron coordinates. The above equation holds for each of these coordinates. Thus, in summary:
This interpretation explains for example the linear relationship of the oscillator strength of the lowest excited state with system size in the case of some conjugated organic polymers (see e.g. this paper): If there are more electrons available to oscillate, then the transition strength increases.
The oscillator strength fij between two non-degenerate states i and j is defined (in atomic units) as two thirds of the squared transition dipole moment multiplied by the energy gap
where the vector r contains all 3N spatial coordinates of the N electronsThe Thomas-Reiche-Kuhn sum rule now states that the sum over the oscillator strengths from one state i to all possible other states is equal to the number of electrons in the system, i.e.
In particular, if we consider excitations from the ground state, then all oscillator strengths are positive. Which means that the oscillator strengths can in fact be viewed as a partitioning of the number of electrons.
The derivation of this sum rule starts by realizing that the momentum operator with respect to any spatial coordinate x of any particle (e.g. x=y2) is given as the commutator of the Hamiltonian with this coordinate
This follows whenever H is of the form
where clearly the derivatives with respect to x are the only part, which does not commute with x itself
By applying the product rule twice, the first term of this expression becomes
And in summary
The remaining proof follows what is shown here (sorry that I am switching the notation, but I copy-and-pasted a little bit ...). First one realizes that the commutator of x and px is equal to i
Then one expands the commutators and inserts a resolution of the identity over the eigenstates of the Hamiltonian
Insert the above expression for px
The commutators are evaluated by letting H act either on the bra or the ket, which results in a multiplication with the respective eigenvalue. And after summing together the equivalent terms one obtains
The actual r vector was composed of 3N individual electron coordinates. The above equation holds for each of these coordinates. Thus, in summary:
which is just what we wanted to show.
Edit: The discussion is also related to the relation of the length and velocity gauge for computing oscillator strengths. Some more information here.
Saturday, 2 November 2013
Phosphorescent Complexes
If you are into phosphorescent transition metal complexes here is a paper for you: The triplet state of organo-transition metal compounds. Triplet harvesting and singlet harvesting for efficient OLEDs by Yersin et al. It was very interesting to read this. They just give a relly nice picture of spin-orbit coupling effects and how they are affected by the ligand field. As a major effect: if the ligand field allows for quasi-degenerate states (e.g. octahedral complexes), then the zero-field splitting will be much stronger than in other cases (e.g square planar). And later on they show that the zero-field splitting shows a strong inverse correlation with the phosphoresence lifetime. But on the other hand, if the zero field splitting is too strong, then the highest triplet substate will no longer be efficiently thermally populated, which is a problem since this is usually the state with the strongest emission strength. For this reason there is as of yet no triplet harvesting complex with a phosphorescent lifetime below 1 μs.
Of course, I do not know if it is all correct. But it is nice to see that people are actually taking their time to produce realistic models to explain their results.
Of course, I do not know if it is all correct. But it is nice to see that people are actually taking their time to produce realistic models to explain their results.
Sunday, 23 June 2013
DNA excimers
Here is our new paper: "Electronic excitation and structural relaxation of the adenine dinucleotide in gas phase and solution." In this article we looked at excited state relaxation and excimer formation between two stacked adenine molecules.
The new result of this study is that orbital interactions might play an important role that is often overlooked when excited state interactions are only considered in the framework of charge transfer or Frenkel excitonic states (similar to what we found for the naphthalene excimer [JCTC 2012, 137, 22A514] ). And the force holding together the excimer may even have some parallels to chemical bonding. In the case of pyrimidine bases, actual bonds may be formed and dimerization occurs. This is of course not the case for adenine. But our results suggest that it is possible to interpret the excimer state between adenines in a similar framework only that (fortunately for us) no stable dimer can be formed.
Of course all computational results are speculative in such an area as you do not only need an accurate description of the excited states but also of the intermolecular interaction potential and the environment, which are difficult tasks even when considered by itself. So I am not claiming that this is the final solution. But I hope it is a step in the right direction.
The new result of this study is that orbital interactions might play an important role that is often overlooked when excited state interactions are only considered in the framework of charge transfer or Frenkel excitonic states (similar to what we found for the naphthalene excimer [JCTC 2012, 137, 22A514] ). And the force holding together the excimer may even have some parallels to chemical bonding. In the case of pyrimidine bases, actual bonds may be formed and dimerization occurs. This is of course not the case for adenine. But our results suggest that it is possible to interpret the excimer state between adenines in a similar framework only that (fortunately for us) no stable dimer can be formed.
Of course all computational results are speculative in such an area as you do not only need an accurate description of the excited states but also of the intermolecular interaction potential and the environment, which are difficult tasks even when considered by itself. So I am not claiming that this is the final solution. But I hope it is a step in the right direction.
Thursday, 26 July 2012
UV absorption in DNA
There is another paper by us, which is just in the final stages of being published: "The UV absorption spectrum of alternating DNA duplexes - analysis of excitonic and charge transfer interactions". The idea was to simulate the absorption spectra of alternating DNA duplexes and analyze the excited states according to charge transfer character and excitonic delocalization.
The method we used was ADC(2), which has the advantage of not being as computationally expensive as CASPT2 while not being TDDFT... By now also the TDDFT studies have rather convergent results and raised the charge transfer states to the energy of the bright states. But it took a lot of reparametrization and "alchemy" to get there in my opinion. But then I am of course not an expert when it comes to TDDFT ...
The environment was simulated by QM/MM electrostatic embedding. Advantage: atomistic description of the environment, disadvantage: neglect of all non-electrostatic terms. The spectra decomposition was performed according to the analysis method described in this post.
The main results are
The method we used was ADC(2), which has the advantage of not being as computationally expensive as CASPT2 while not being TDDFT... By now also the TDDFT studies have rather convergent results and raised the charge transfer states to the energy of the bright states. But it took a lot of reparametrization and "alchemy" to get there in my opinion. But then I am of course not an expert when it comes to TDDFT ...
The environment was simulated by QM/MM electrostatic embedding. Advantage: atomistic description of the environment, disadvantage: neglect of all non-electrostatic terms. The spectra decomposition was performed according to the analysis method described in this post.
The main results are
- the states are rather localized
- charge transfer states are at higher energies than the bright states in the Franck-Condon region
- there is a significant coupling between charge transfer and locally excited (Frenkel) states
- spectral broadening is mainly caused by intramolecular vibrations
Friday, 22 June 2012
Excitons and charge transfer states
Our new paper was just accepted by the Journal of Chemical Theory and Computation. And it is actually already available as a just accepted manuscript (Link). What we are looking at is the analysis of excited states. If an excited state contains several configurations, it is not only tedious to analyze it but a lot of information may be missed by considering the configurations independently. The problem is in particular significant in the case of delocalized states. Here our procedure, which is based on the transition density matrix, can provide a well defined automatized measure for delocalization, which is also independent on the orbital resolution.
Another challenging case is to differentiate between (Frenkel) excitonic and charge resonance transitions. For example if you look at a symmetric dimer: Then you know that there have to be excitonic resonance states and charge resonance states. But because all the orbitals are delocalized, it is not trivial to differentiate between them (unless you use our analysis tools).
In summary, I am kind of proud of it ... :) And interestingly one of my first blog posts about quantum chemistry, which I have not really thought of in the meantime, was concerned with a part of this paper: a singular value decomposition of the transition density matrix yielding the natural transition orbitals. Apparently I had the same taste in things I consider interesting five years ago...
Another challenging case is to differentiate between (Frenkel) excitonic and charge resonance transitions. For example if you look at a symmetric dimer: Then you know that there have to be excitonic resonance states and charge resonance states. But because all the orbitals are delocalized, it is not trivial to differentiate between them (unless you use our analysis tools).
In summary, I am kind of proud of it ... :) And interestingly one of my first blog posts about quantum chemistry, which I have not really thought of in the meantime, was concerned with a part of this paper: a singular value decomposition of the transition density matrix yielding the natural transition orbitals. Apparently I had the same taste in things I consider interesting five years ago...
Thursday, 19 April 2012
Excited state proton transfer in solution
If you are interested in excited state proton transfer, you can check out our new paper: "The effect of hydrogen bonding on the excited-state proton transfer in 2-(2′-hydroxyphenyl)benzothiazole: a TDDFT molecular dynamics study"

The idea was to follow the excited state proton transfer through the water network by dynamics simulations. And actually what was found is that there are two possibilities: either a direct intramolecular proton transfer, which is known from aprotic solvents; but also proton transfer through the network.
The work was down by Nawee Kungwan who was a guest at our institute some time ago. My involvement was mostly through technical support and discussions, and by doing some early test calculations.
Sunday, 18 March 2012
Rydberg states
Rydberg states can be seen as ionized molecules that are circled by a loosely bound electron. Because of their diffuse character, the central molecule appears almost like a point charge to the electron. And the states resemble the excited states of atomic hydrogen (the Rydberg series).
The example we are currently working on is morpholine. The highest occupied molecular orbital is a regular tight non-bonding orbital.
 The LUMO (which also corresponds to the first excited state) is a diffuse s-type orbital.
The LUMO (which also corresponds to the first excited state) is a diffuse s-type orbital.
 The next three orbitals (yielding the next three states) are of p-type character. The first one is antisymmetric with respect to the molecular symmetry plane (i.e. it is of a" character).
The next three orbitals (yielding the next three states) are of p-type character. The first one is antisymmetric with respect to the molecular symmetry plane (i.e. it is of a" character).
 And two more p-type orbitals follow in the other spatial directions.
And two more p-type orbitals follow in the other spatial directions.


It took some attempts to actually get nice pictures of the Rydberg orbitals. The problem is that they are very large and most programs in their default settings will not sample these areas. What I ended up doing is the following:
Use Turbomole to create
The example we are currently working on is morpholine. The highest occupied molecular orbital is a regular tight non-bonding orbital.
 The LUMO (which also corresponds to the first excited state) is a diffuse s-type orbital.
The LUMO (which also corresponds to the first excited state) is a diffuse s-type orbital. The next three orbitals (yielding the next three states) are of p-type character. The first one is antisymmetric with respect to the molecular symmetry plane (i.e. it is of a" character).
The next three orbitals (yielding the next three states) are of p-type character. The first one is antisymmetric with respect to the molecular symmetry plane (i.e. it is of a" character). And two more p-type orbitals follow in the other spatial directions.
And two more p-type orbitals follow in the other spatial directions.

It took some attempts to actually get nice pictures of the Rydberg orbitals. The problem is that they are very large and most programs in their default settings will not sample these areas. What I ended up doing is the following:
Use Turbomole to create
.plt files but use a non-standard larger grid.$pointval mo 23-30
grid1 vector 1 0 0 range -15,15 points 200
grid2 vector 0 1 0 range -15,15 points 200
grid3 vector 0 0 1 range -15,15 points 200
Wednesday, 8 April 2009
No Fluorescence (2)
So how is non-radiative decay possible? I showed the conical intersections of cytosine some time ago. These geometries, where the first excited and ground state are of the same energy, exist. But the chance of exactly reaching one is zero (since they are only an N-2 dimensional hyperline in the N-dimensional space of geometries of a molecule).
The point is that the picture of isolated electronic states breaks down, i.e. the Born-Oppenheimer approximation. In solid state chemistry that is what they call electron-phonon-coupling.
If you consider the nuclei in terms of an external perturbation, you can take at look at the adiabatic theorem: If nuclear motion is fast and there are close lying electronic states, the states will mix. You get the fast nuclear motion from the excitation energy. About close lying states: Typically all the excited states are close and the molecule quickly reaches S1 if it was initially excited to a higher state (Kasha's rule). If there are suitable intersections to the ground state it can even relax completely.
For doing molecular dynamics you need the energy of your electronic state Ψi as always. But you also need couplings to the other states:
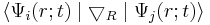
They are called the non-adiabatic coupling vectors and give the major post Born-Oppenheimer contributions.
Here is a little dynamics movie. This is a test MCSCF run on cytosine. The electronic structure is from the Columbus program and the non-adiabatic surface-hopping dynamics are with Newton-X (both from our group actually).
The idea of surface-hopping is that we still want to have a classical trajectory as a basis, but post Born-Oppenheimer corrections are introduced through jumping in between states. Here you can see the cytosine molecule with color coded electronic states: green S1, orange S2. You see how the molecule jumps into the second excited state intermediately and then stays in the first excited state. There is a lot of motion because of the high excess energy.
There is no decay to the ground state in these 341 fs simulated. It should happen soon after or tanning would be extremely dangerous. But the time simulated is still rather short, it is for example a ten-thousandth of the typical fluorescence life-time.
The point is that the picture of isolated electronic states breaks down, i.e. the Born-Oppenheimer approximation. In solid state chemistry that is what they call electron-phonon-coupling.
If you consider the nuclei in terms of an external perturbation, you can take at look at the adiabatic theorem: If nuclear motion is fast and there are close lying electronic states, the states will mix. You get the fast nuclear motion from the excitation energy. About close lying states: Typically all the excited states are close and the molecule quickly reaches S1 if it was initially excited to a higher state (Kasha's rule). If there are suitable intersections to the ground state it can even relax completely.
For doing molecular dynamics you need the energy of your electronic state Ψi as always. But you also need couplings to the other states:
They are called the non-adiabatic coupling vectors and give the major post Born-Oppenheimer contributions.
Here is a little dynamics movie. This is a test MCSCF run on cytosine. The electronic structure is from the Columbus program and the non-adiabatic surface-hopping dynamics are with Newton-X (both from our group actually).
The idea of surface-hopping is that we still want to have a classical trajectory as a basis, but post Born-Oppenheimer corrections are introduced through jumping in between states. Here you can see the cytosine molecule with color coded electronic states: green S1, orange S2. You see how the molecule jumps into the second excited state intermediately and then stays in the first excited state. There is a lot of motion because of the high excess energy.
There is no decay to the ground state in these 341 fs simulated. It should happen soon after or tanning would be extremely dangerous. But the time simulated is still rather short, it is for example a ten-thousandth of the typical fluorescence life-time.
Friday, 6 July 2007
Retinal - Ultrafast is just about fast enough
Ultrafast, barrierless processes have the remarkable feature that they occur in less than a picosecond and can therefore be modelled with molecular dynamics. The cis-trans isomerisation of photo-excited retinal is ultrafast. This is not primarily God's grace toward theoretical chemists (who like to model the process) but it ensures that the process is efficient and vision works. [1]
The trick with an ultrafast process is that after excitation, the reaction occurs (almost) without a barrier. The time is determined by the skeletal deformations, cf. a 333/cm vibration has a period of 100 fs.
To understand what is going on, it is sufficient to consider a smaller system than retinal, for example the pentadiene-iminium ion (the N instead of O comes from the protein). The system is strongly polarised toward the N. The first excited state (shwon in red) is polarized the opposite way.
 After excitation (γ means "photon" [2]) the cis-doublebond becomes a single bond. Because of that it stretches out. Now there is an ecliptical single bond with weak π-conjugation. It spontaniously rotates.
After excitation (γ means "photon" [2]) the cis-doublebond becomes a single bond. Because of that it stretches out. Now there is an ecliptical single bond with weak π-conjugation. It spontaniously rotates.
The crucial step comes now: The excited state decreases in energy as the rotation progresses. The ground state's energy increases. At about 90° torsion angle their energies are the same. This is called a conical intersection. Now a radiationless transition occurs and we are in the ground state again. Ground state means double bond. And that means that a linear alignment is preferred. The molecule can either move back to the cis form or it keeps rotating the same direction and becomes trans. The ratio depends on the exact shape of the potential energy surface.
During that whole process we move down on potential energy surfaces. The geometry is relaxed to a minimum in the excited state. Then the molecule switches to the ground state where it has a local maximum. And the reaction keeps going until we reach one of the stable ground state structures. The energy needed to drive the process comes from the photon. But as soon as the photon is absorbed everything happens spontaneously.
Because of the short reaction time no competing reactions are to be expected. Fluorescence is with 10-8 seconds 5 orders of magnitude slower. But also collisions that could take up energy aren't likely in that kind of a time frame. That means we can expect high quantum yield. It is 65% for retinal in rhodopsin and 25% for retinal in solution.
The information was mostly from this article by Garavelli (10.1007/s00214-005-0030-z).
[1] Sorry for making a religious reference in a scientific text. But where I come from the gap between religion and science isn't quite as big. Our religious people don't try to sabotage scientific theories for incomprehensible reasons and our scientists don't announce that they are atheists all the time. I don't know whose fault the situation in the US is. Maybe religious people should be nice and tolerant. Maybe scientists should be smart and on top of things. Maybe it's difficult for anyone to stop a fight once it has that scale and all you can do is defend your own interests.
[2] I don't like the hν for two reasons. First the energy is neither the only feature of a photon (it has momentum, spin, ...) nor is hν a unique identification for it. Second I don't like ν because it looks like a v and it does not abbreviate the word "frequency" like f does.
The trick with an ultrafast process is that after excitation, the reaction occurs (almost) without a barrier. The time is determined by the skeletal deformations, cf. a 333/cm vibration has a period of 100 fs.
To understand what is going on, it is sufficient to consider a smaller system than retinal, for example the pentadiene-iminium ion (the N instead of O comes from the protein). The system is strongly polarised toward the N. The first excited state (shwon in red) is polarized the opposite way.
 After excitation (γ means "photon" [2]) the cis-doublebond becomes a single bond. Because of that it stretches out. Now there is an ecliptical single bond with weak π-conjugation. It spontaniously rotates.
After excitation (γ means "photon" [2]) the cis-doublebond becomes a single bond. Because of that it stretches out. Now there is an ecliptical single bond with weak π-conjugation. It spontaniously rotates.The crucial step comes now: The excited state decreases in energy as the rotation progresses. The ground state's energy increases. At about 90° torsion angle their energies are the same. This is called a conical intersection. Now a radiationless transition occurs and we are in the ground state again. Ground state means double bond. And that means that a linear alignment is preferred. The molecule can either move back to the cis form or it keeps rotating the same direction and becomes trans. The ratio depends on the exact shape of the potential energy surface.
During that whole process we move down on potential energy surfaces. The geometry is relaxed to a minimum in the excited state. Then the molecule switches to the ground state where it has a local maximum. And the reaction keeps going until we reach one of the stable ground state structures. The energy needed to drive the process comes from the photon. But as soon as the photon is absorbed everything happens spontaneously.
Because of the short reaction time no competing reactions are to be expected. Fluorescence is with 10-8 seconds 5 orders of magnitude slower. But also collisions that could take up energy aren't likely in that kind of a time frame. That means we can expect high quantum yield. It is 65% for retinal in rhodopsin and 25% for retinal in solution.
The information was mostly from this article by Garavelli (10.1007/s00214-005-0030-z).
[1] Sorry for making a religious reference in a scientific text. But where I come from the gap between religion and science isn't quite as big. Our religious people don't try to sabotage scientific theories for incomprehensible reasons and our scientists don't announce that they are atheists all the time. I don't know whose fault the situation in the US is. Maybe religious people should be nice and tolerant. Maybe scientists should be smart and on top of things. Maybe it's difficult for anyone to stop a fight once it has that scale and all you can do is defend your own interests.
[2] I don't like the hν for two reasons. First the energy is neither the only feature of a photon (it has momentum, spin, ...) nor is hν a unique identification for it. Second I don't like ν because it looks like a v and it does not abbreviate the word "frequency" like f does.
Wednesday, 4 July 2007
Rhodopsin
The interesting thing about retinal is its photochemistry. I will talk about that soon. But this is another one of the protein-with-functional-group-picture posts. I still haven't quite realized how nice PyMOL is and I still enjoy making pictures with it. But of course also thanks to Röntgen and Bragg. And Edwards et al. who investigated the structure of this bovine rhodopsin.
This is the subunit with retinal (shown in red). Long stretched helices.
 Retinal is bonded to a lysine residue (pink) to form an iminium ion. Or protonated Schiff base as they say if they are trying to confuse me. Retinal fits nicely into the helices. The π-system is almost planar only the double bond in the ring kind of sticks out. The 11-cis-bond (where I don't understand why it is called 11, probably a terpene nomenclature) is behind the green arc.
Retinal is bonded to a lysine residue (pink) to form an iminium ion. Or protonated Schiff base as they say if they are trying to confuse me. Retinal fits nicely into the helices. The π-system is almost planar only the double bond in the ring kind of sticks out. The 11-cis-bond (where I don't understand why it is called 11, probably a terpene nomenclature) is behind the green arc.
 Carboxylic groups surrounding retinal are important for the charge distribution. You can select them in PyMOL with the following command (after making a selection "retinal" with retinal):
Carboxylic groups surrounding retinal are important for the charge distribution. You can select them in PyMOL with the following command (after making a selection "retinal" with retinal):
There is one at the iminium group ...
 ... and one at the β-ionone ring.
... and one at the β-ionone ring.
 They give the molecule kind of a jungle feeling.
They give the molecule kind of a jungle feeling.
This is the subunit with retinal (shown in red). Long stretched helices.
 Retinal is bonded to a lysine residue (pink) to form an iminium ion. Or protonated Schiff base as they say if they are trying to confuse me. Retinal fits nicely into the helices. The π-system is almost planar only the double bond in the ring kind of sticks out. The 11-cis-bond (where I don't understand why it is called 11, probably a terpene nomenclature) is behind the green arc.
Retinal is bonded to a lysine residue (pink) to form an iminium ion. Or protonated Schiff base as they say if they are trying to confuse me. Retinal fits nicely into the helices. The π-system is almost planar only the double bond in the ring kind of sticks out. The 11-cis-bond (where I don't understand why it is called 11, probably a terpene nomenclature) is behind the green arc. Carboxylic groups surrounding retinal are important for the charge distribution. You can select them in PyMOL with the following command (after making a selection "retinal" with retinal):
Carboxylic groups surrounding retinal are important for the charge distribution. You can select them in PyMOL with the following command (after making a selection "retinal" with retinal):select SurrAc, (retinal around 10) and resn asp+gluThere is one at the iminium group ...
 ... and one at the β-ionone ring.
... and one at the β-ionone ring. They give the molecule kind of a jungle feeling.
They give the molecule kind of a jungle feeling.
Subscribe to:
Posts (Atom)