Sunday, 23 June 2013
DNA excimers
The new result of this study is that orbital interactions might play an important role that is often overlooked when excited state interactions are only considered in the framework of charge transfer or Frenkel excitonic states (similar to what we found for the naphthalene excimer [JCTC 2012, 137, 22A514] ). And the force holding together the excimer may even have some parallels to chemical bonding. In the case of pyrimidine bases, actual bonds may be formed and dimerization occurs. This is of course not the case for adenine. But our results suggest that it is possible to interpret the excimer state between adenines in a similar framework only that (fortunately for us) no stable dimer can be formed.
Of course all computational results are speculative in such an area as you do not only need an accurate description of the excited states but also of the intermolecular interaction potential and the environment, which are difficult tasks even when considered by itself. So I am not claiming that this is the final solution. But I hope it is a step in the right direction.
Thursday, 26 July 2012
UV absorption in DNA
The method we used was ADC(2), which has the advantage of not being as computationally expensive as CASPT2 while not being TDDFT... By now also the TDDFT studies have rather convergent results and raised the charge transfer states to the energy of the bright states. But it took a lot of reparametrization and "alchemy" to get there in my opinion. But then I am of course not an expert when it comes to TDDFT ...
The environment was simulated by QM/MM electrostatic embedding. Advantage: atomistic description of the environment, disadvantage: neglect of all non-electrostatic terms. The spectra decomposition was performed according to the analysis method described in this post.
The main results are
- the states are rather localized
- charge transfer states are at higher energies than the bright states in the Franck-Condon region
- there is a significant coupling between charge transfer and locally excited (Frenkel) states
- spectral broadening is mainly caused by intramolecular vibrations
Wednesday, 12 October 2011
Orbitals, example
What I am showing are the frontier orbitals of an adenine-thymine stack inside of DNA. The calculations were QM/MM calculations where the eletrons of the two bases were treated quantum mechanically and the remaining DNA and waters were considered as point charges. The orbitals are shown in order of decreasing energy with a qualitative label below them.

Thymine - π*

Adenine - π* (LUMO)

Adenine - π (HOMO)

Thymine - π

Adenine - π

Adenine - π

Adenine - n

Thymine - π

Thymine - n
Monday, 4 April 2011
DNA base pairing
First a look at the adenine-thymine base pair. Thymine is on the strand whose 5' phosphate group is shown on top. The strand with adenine goes the opposite way and you see an open 3' oxygen. There are two hydrogen bonds in the AT pair.

Let's look at this base pair from the other side. The important point is that the backbone looks the same as in the above image. And ths is why you can have alternating A's and T's in a strand. Apparently this is what Francis Crick pointed out to James Watson, when Watson had suggested base pairing. Considering that this had to be pointed out even to the person winning the Nobel prize for discovering the DNA structure, I think it is safe to say that people without this understanding of helical symmetry do not have to feel bad about it. But apparently it is a non-trivial fact.

The next non-trivial fact is that the backbone of the guanine-cytosine pair looks just like the AT and TA pairs. Here you can see the base pair with its three hydrogen bonds and which is thereore bonded a little bit more strongly.

And again, CG also fits.

It seems therefore that one reason for the specific choice of bases is that they all fit into the same helical structure. And therefore it is possible to have a stable helix which contains the genetic code through alternating bases. Another source of selection pressure was probably photostability, as explained here.
Tuesday, 29 March 2011
DNA close up
I wanted to look a little bit more at the details of the DNA structure. In part motivated by James Watson's book (described here). Some time ago I showed pictures of the helix, but now I want to show some molecular details.[1]
This is the backbone of DNA in its B-form. The phosphate is in front. The 2-deoxyribose is the vertical 5-ring. Its anomeric center is bonded to the nitrogen of the base and the oxygen closing the furanose ring. The 3' oxygen bonds to the next phosphate group. The 5' oxygen leading to the other side is bonded to the carbon outside of the ring.

In principle a pentose has five active groups. In DNA all are saturated and no OH groups remain. Why?
1' bonded to the nitrogen
2' that's the "deoxy" in "deoxy-ribonucleic acid"
3' bonded to phosphate
4' forming the ring
5' bonded to phosphate
What happens if we put an OH group at the 2' position?

The new OH groups is shown with its full van-der-Waals radius. There is clearly not enough room for the extra oxygen atom, considering for example that the other oxygen atoms should be just as big. It is in fact not possible to push the atoms out of the way in any simple manner. Therefore RNA does not form a B-helix.
The structures were created with the nucleic functionality of Tinker. The pictures were drawn with pymol.
[1] One thing I really learned in science is quoting myself ...
Wednesday, 1 December 2010
DNA photostability

The images correspond to minimum energy conical intersection structures of the four DNA bases. At such geometries the ground state energy is strongly raised but the energy of the biradical excited state is not so much affected or even lowered. Therefore ground and excited states can become degenerate and a transition can occur. It is probably through such geometric distortions that DNA bases can efficiently relax to the ground state on a pico second time scale. Then only a slight chance of undergoing any photochemical modifications remains.
Through this inherent photostability in can be assured that only a small fraction of DNA bases undergoes photodamage. The small part that does react, for example to pyrimidine dimers, can be repaired by enzymes.
Now the technical part about the image: The background comes from a picture of the sun against a blue sky in the Austrian alps. The molecular structures were rendered with pymol. The trick was to use a transparent background in the rendering
set ray_opaque_background=0Through adjusting the light setting I tried to get the light source to where the sun would be for the separate structures. e.g.
set light, [-100,-20,0]to move the light source to the left and a little bit lower which would have been the appropriate setting for the adenine on top.
By reducing the
ambient setting I tried to make this more pronounced. For this I used the Setting/Edit All in the menu, which is in some cases more convenient than the command line.
Monday, 11 May 2009
DNA
 Or with thicker sticks:
Or with thicker sticks:
In fact there is of course not so much space between the bases. Here is what it looks like when you draw the van der Waals spheres for the bases. It is actually quite difficult to model this stacking interaction because it is almost pure dispersion. From a computational point of view, dispersion is electron correlation. If the effect you are interested in is pure electron correlation, then you'd better model it really well. Hartree-Fock gives you zero dispersion. MP2 is great if you kind of want to include some electron correlation but it is not accurate enough in this case. What they are actually doing is high level coupled cluster CCSD(T) extrapolated to the complete basis set limit.

Another interesting question is how defects are propagated in such a framework: holes, electrons, or electron hole pairs. First of course you want to preserve the integrity of your genome. And second you want to make nano-robots. DNA is already a self-assembling structure with molecular recognition. If charge transport is better understood and conductive analogues are found, DNA will kick nanotubes' ass.
By the way: for printing out pymol graphics (see again [1]), the ray_trace_mode setting is nice. Especially:
set ray_trace_mode=1Then you get some nice black frames instead of fuzzi ends in the print.

[1] from the crystal structure 1BNA. And drawn with pymol. Maybe I should eventually switch from pymol because they are becoming more and more commercial. But then I guess they would not mind me as a little personal user.
Monday, 24 November 2008
Ribonuclease A
Ribonoclease A is a sturdy little molecule with 124 amino acids. Its four disulfide bridges (yellow in the picture) make it very stable and comparably easy to isolate. Just 124 amino acids also makes it nice for computations. The pictures were made with VMD which makes nice pictures and is also a pretty versatile tool that reads tons of file formats.

The active group of Ribonuclease A comes from two Histidine (black) residues that help hydrolizing the phosphate in the nucleic acid. Lysine (orange) provides the positive charge to interact with negatively charged phosphate groups.
 The first thing you can do with a molecular structure is an energy minimization. These are the Cα traces of the crystal structure (yellow) and the vacuum optimized structure (red). They are in fact quite different even though the red structure is just the next local minimum.
The first thing you can do with a molecular structure is an energy minimization. These are the Cα traces of the crystal structure (yellow) and the vacuum optimized structure (red). They are in fact quite different even though the red structure is just the next local minimum. The major problem with this optimization is that I did not include solvent effects. The electrostatic interaction energy (in the macroscopic limit) is given by
The major problem with this optimization is that I did not include solvent effects. The electrostatic interaction energy (in the macroscopic limit) is given byAnother thing you can do with structures is normal mode analysis. That means computing the eigenvalues/eigenvectors of the mass weighted energy Hessian matrix which correspond to the frequencies/motions of the normal modes. Ribonuclease A is small for a protein but it still has 1856 atoms. So we have 3x1856=5568 degrees of freedom. We have to compute, store and process about 30 million matrix elements. This seems much but these days most regular computers will be able to do this in less than half an hour. 30 million matrix elements corresponds to something like 300 MByte, nothing too crazy. [2]
These are the three lowest frequency modes (aside from translation and rotation) of Ribonuclease A. They are large scale backbone motions. A weak force constant and a high effective mass causes them to have wave numbers below 10/cm.



[1] Not the kind of lab where you need a lab coat.
[2] Actually if you are just interested in a few eigenvalues and -vectors, you can go much higher. For example multi-reference configuration interaction with Columbus (which is a great program by the way....): finding the first 4 states out of a 8569890 x 8569890 matrix was possible within about 17 hours. In this case you don't store trillions (7E13) of matrix elements and do a diagonalization. You use the iterative Davidson procedure which will find an eigenvector close to a starting guess.
Sunday, 9 November 2008
GFP pictures
In the sphere model all proteins pretty much look alike. But I still think kind of cool, especially if you use Lightnir's QuteMol preset for pymol.
 The cartoon model shows you more about the structure. GFP consists of a β-sheet "barrel" that encapsulates the chromophore. You need a rigid structure if you want to make sure that the chromophore does not quench before it is able to show fluorescence. It is interesting to compare this to Rhodopsin which is apparently much more flexible. This makes sense since in Rhodopsin the biological activity is related to non-radiative decay and isomerisation.
The cartoon model shows you more about the structure. GFP consists of a β-sheet "barrel" that encapsulates the chromophore. You need a rigid structure if you want to make sure that the chromophore does not quench before it is able to show fluorescence. It is interesting to compare this to Rhodopsin which is apparently much more flexible. This makes sense since in Rhodopsin the biological activity is related to non-radiative decay and isomerisation. The chromophore is trapped at the center of the barrel.
The chromophore is trapped at the center of the barrel.
Interestingly the chromophore is made directly out of amino acids. You can take a look at it and think of how this is done.
 The answer is here. The hydroxyphenyl comes from tyrosine (as expected). The imidazolon ring is formed after cyclization, the bridging double bond through oxidation. It is interesting to consider that this apparently works in many different organisms and not just in the jelly fish aequoria victoria. The driving force seems to be that the amino acids are pressed together in the barrel.
The answer is here. The hydroxyphenyl comes from tyrosine (as expected). The imidazolon ring is formed after cyclization, the bridging double bond through oxidation. It is interesting to consider that this apparently works in many different organisms and not just in the jelly fish aequoria victoria. The driving force seems to be that the amino acids are pressed together in the barrel.The chromophore is part of the protein chain. It is connected to the helix that goes through the center of the protein.
 Or from a different perspective.
Or from a different perspective.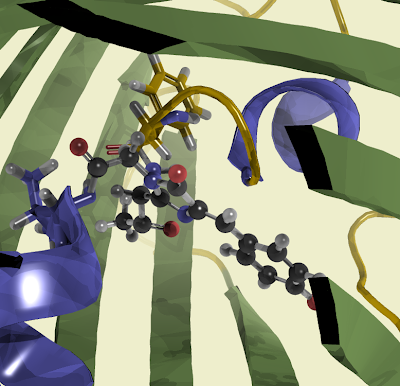
Sunday, 7 September 2008
fold it
This what it looks like. Notice the amazing score of 9577 for this Calcium Ion binding protein. (I am leaving a little head start to lightnir ...)

The producers of the game mention that the computer game strategy may even be a fruitful way of tackling protein folding, which is basically the attempt to find the global free energy minimum structure of a protein. For every geometry you can compute the energy and forces. You can let the forces pull the geometry toward a local energy minimum. But the energy will usually be a very complex function of the geometry with many local minima. And this local minimum may be far away from the global minimum.
One strategy to go on is dynamics. Simulate protein motion at some finite temperature and hope that it will eventually overcome the barrier, leave the minimum, and go to a lower energy minimum.
The second strategy is Monte Carlo. Trying to systematically improve the structure by applying random changes.
The third strategy is making a game out of it and telling people it's fun.
Another interesting trivia about protein folding: Folding a random co-polymer is an NP-complete problem, meaning it has exponential scaling which makes it impossible to use for all but the smallest systems. The reason why in silico protein folding still works in some cases is first that we know the building blocks well and have a big knowledge base. The second reason is that proteins are folding in nature so we should be able to emulate this process and that there is a selection pressure for proteins to fold easily, so it works in nature.
The last thing to wonder about is the question between computer and human problem solving abilities. It will take something like 50 years until we can build something with the processing power of even an ant. Still we are using computers rather than ants for our everyday problems. Well it's because computers are more flexible. But it seems with anything that is remotely related to anything that evolution would select us to do we kick their asses.
Sunday, 8 June 2008
Glucocorticoid receptor (2)
You can take a better lock at the ligand inside the binding pocket.

Or you can make this picture which looks kind of cool, I think.

This one shows the opening in the outside surface.

Two commands to mention are:
Changing the transparency of an object ("cha"):
set transparency, .1, chaI can never quite remember the syntax for that.
Pymol has a nice way of parallelizing the rendering. The downside is that this will stop the whole system. If you want to keep working while rendering in a multicore machine, you can change the maximum threads:
set max_threads, 1
Sunday, 1 June 2008
Glucocorticoid receptor
The whole protein looks like this, the ligand is shown in red.
 If you draw the protein with its van der Waals surface, you notice that there is a small opening which is probably the place where the ligand entered the binding pocket.
If you draw the protein with its van der Waals surface, you notice that there is a small opening which is probably the place where the ligand entered the binding pocket.
Apparently the receptor closed again and the molecule is trapped in there.

If you eat away some of the outer amino acids, you can take a better look at the binding pocket.

Or at a semi transparent binding pocket:
 Remove some more amino acids that are in the way and it'll look like this.
Remove some more amino acids that are in the way and it'll look like this.
How do you create those picutres in pymol?
First you select one chain (the fact that there are four chains is just because that's how it crystallizes):
select cha, chain aIf you go for "preset pretty", you can see the ligand and select it. Call it lig. Then you can modify it, give it double bonds, show it as ball and stick model, ...
For the main protein you may want to select the different secondary structures:
select helix, ss h and cha2
select sheet, ss s and cha2
select loop, cha2 and not (ss h or ss s)Then you can color those the way you want.
If you want a surface, it makes sense to duplicate that chain a and show the duplicate as a surface.
If you want to select the binding pocket, go for:
select pocket, (lig around 5)Then you can erase some of the amino acids that are in the way.
Of course after doing all this, you have to play around with colors, perspectives and rendering methods.
Sunday, 4 May 2008
tRNA
The secondary structure looks like this, the clover leaf. The amino acid is bound at the top, the anticodon at the bottom. The lines represent the base pairing.

For the 3D structure you have to first consider that helices are formed with base pairing - twist around all those parts. In tRNA there is an important tertiary element, the connection between the left and right loop - fold the molecule. You will get something like this.

The open end is shown in read, the anticodon green, the right loop yellow, the left loop cyan.
Twisted around:
 The open end:
The open end: The anticodon loop:
The anticodon loop:
These three bases are probably the anticodon. They are GAA which correctly codes for Phe which this piece of tRNA is supposed to do.

If you want to move the molecule yourself you can find the link to a jmol applet from the pdb.

